Data preparation
Now the positive mode and negative mode have been annotated respectively. We need to merge positive and negative mode data.
library(tidymass)
#> Registered S3 method overwritten by 'Hmisc':
#> method from
#> vcov.default fit.models
#> ── Attaching packages ──────────────────────────────────────── tidymass 1.0.9 ──
#> ✔ massdataset 1.0.34 ✔ metid 1.2.34
#> ✔ massprocesser 1.0.10 ✔ masstools 1.0.13
#> ✔ masscleaner 1.0.12 ✔ dplyr 1.1.4
#> ✔ massqc 1.0.7 ✔ ggplot2 3.5.1
#> ✔ massstat 1.0.6 ✔ magrittr 2.0.3
#> ✔ metpath 1.0.8
library(tidyverse)
#> ── Attaching core tidyverse packages ──────────────────────── tidyverse 2.0.0 ──
#> ✔ forcats 1.0.0 ✔ readr 2.1.5
#> ✔ lubridate 1.9.3 ✔ stringr 1.5.1
#> ✔ purrr 1.0.2 ✔ tibble 3.2.1
#> ── Conflicts ────────────────────────────────────────── tidyverse_conflicts() ──
#> ✖ xcms::collect() masks dplyr::collect()
#> ✖ MSnbase::combine() masks Biobase::combine(), BiocGenerics::combine(), dplyr::combine()
#> ✖ tidyr::expand() masks S4Vectors::expand()
#> ✖ tidyr::extract() masks magrittr::extract()
#> ✖ metid::filter() masks metpath::filter(), dplyr::filter(), massdataset::filter(), stats::filter()
#> ✖ S4Vectors::first() masks dplyr::first()
#> ✖ xcms::groups() masks dplyr::groups()
#> ✖ dplyr::lag() masks stats::lag()
#> ✖ purrr::map() masks mixOmics::map()
#> ✖ BiocGenerics::Position() masks ggplot2::Position(), base::Position()
#> ✖ purrr::reduce() masks MSnbase::reduce()
#> ✖ S4Vectors::rename() masks dplyr::rename(), massdataset::rename()
#> ✖ lubridate::second() masks S4Vectors::second()
#> ✖ lubridate::second<-() masks S4Vectors::second<-()
#> ✖ MASS::select() masks dplyr::select(), massdataset::select()
#> ✖ purrr::set_names() masks magrittr::set_names()
#> ℹ Use the conflicted package (<http://conflicted.r-lib.org/>) to force all conflicts to become errors
Load data
load("metabolite_annotation/object_pos2")
load("metabolite_annotation/object_neg2")
Remove the features without annotations
Positive mode
object_pos2 <-
object_pos2 %>%
activate_mass_dataset(what = "annotation_table") %>%
filter(!is.na(Level)) %>%
filter(Level == 1 | Level == 2)
object_pos2
#> --------------------
#> massdataset version: 0.99.8
#> --------------------
#> 1.expression_data:[ 137 x 259 data.frame]
#> 2.sample_info:[ 259 x 6 data.frame]
#> 259 samples:sample_06 sample_103 sample_11 ... sample_QC_38 sample_QC_39
#> 3.variable_info:[ 137 x 6 data.frame]
#> 137 variables:M86T95_POS M95T100_1_POS M100T160_POS ... M568T622_POS M609T427_POS
#> 4.sample_info_note:[ 6 x 2 data.frame]
#> 5.variable_info_note:[ 6 x 2 data.frame]
#> 6.ms2_data:[ 1042 variables x 951 MS2 spectra]
#> --------------------
#> Processing information
#> 10 processings in total
#> Latest 3 processings show
#> integrate_data ----------
#> Package Function.used Time
#> 1 masscleaner integrate_data() 2024-09-25 19:53:32
#> mutate_ms2 ----------
#> Package Function.used Time
#> 1 massdataset mutate_ms2() 2024-09-25 20:43:24
#> annotate_metabolites ----------
#> Package Function.used Time
#> 1 metid annotate_metabolites_mass_dataset() 2024-09-25 20:44:08.051042
#> 2 metid annotate_metabolites_mass_dataset() 2024-09-25 20:45:30.750549
#> 3 metid annotate_metabolites_mass_dataset() 2024-09-25 20:46:27.89833
Negative mode
object_neg2 <-
object_neg2 %>%
activate_mass_dataset(what = "annotation_table") %>%
filter(!is.na(Level)) %>%
filter(Level == 1 | Level == 2)
object_neg2
#> --------------------
#> massdataset version: 0.99.8
#> --------------------
#> 1.expression_data:[ 125 x 259 data.frame]
#> 2.sample_info:[ 259 x 6 data.frame]
#> 259 samples:sample_06 sample_103 sample_11 ... sample_QC_38 sample_QC_39
#> 3.variable_info:[ 125 x 6 data.frame]
#> 125 variables:M75T52_NEG M89T57_1_NEG M93T204_NEG ... M478T658_NEG M480T641_NEG
#> 4.sample_info_note:[ 6 x 2 data.frame]
#> 5.variable_info_note:[ 6 x 2 data.frame]
#> 6.ms2_data:[ 1092 variables x 988 MS2 spectra]
#> --------------------
#> Processing information
#> 10 processings in total
#> Latest 3 processings show
#> integrate_data ----------
#> Package Function.used Time
#> 1 masscleaner integrate_data() 2024-09-25 19:53:34
#> mutate_ms2 ----------
#> Package Function.used Time
#> 1 massdataset mutate_ms2() 2024-09-25 20:43:34
#> annotate_metabolites ----------
#> Package Function.used Time
#> 1 metid annotate_metabolites_mass_dataset() 2024-09-25 20:47:07.586516
#> 2 metid annotate_metabolites_mass_dataset() 2024-09-25 20:47:44.92965
#> 3 metid annotate_metabolites_mass_dataset() 2024-09-25 20:49:01.646808
Merge data
We need to merge positive and negative mode data as one mass_dataset class.
head(colnames(object_pos2))
#> [1] "sample_06" "sample_103" "sample_11" "sample_112" "sample_117"
#> [6] "sample_12"
head(colnames(object_neg2))
#> [1] "sample_06" "sample_103" "sample_11" "sample_112" "sample_117"
#> [6] "sample_12"
object <-
merge_mass_dataset(x = object_pos2,
y = object_neg2,
sample_direction = "inner",
variable_direction = "full",
sample_by = "sample_id",
variable_by = c("variable_id", "mz", "rt"))
object
#> --------------------
#> massdataset version: 1.0.34
#> --------------------
#> 1.expression_data:[ 262 x 259 data.frame]
#> 2.sample_info:[ 259 x 11 data.frame]
#> 259 samples:sample_06 sample_103 sample_11 ... sample_QC_38 sample_QC_39
#> 3.variable_info:[ 262 x 9 data.frame]
#> 262 variables:M86T95_POS M95T100_1_POS M100T160_POS ... M478T658_NEG M480T641_NEG
#> 4.sample_info_note:[ 11 x 2 data.frame]
#> 5.variable_info_note:[ 9 x 2 data.frame]
#> 6.ms2_data:[ 2134 variables x 1939 MS2 spectra]
#> --------------------
#> Processing information
#> 21 processings in total
#> Latest 3 processings show
#> mutate_ms2 ----------
#> Package Function.used Time
#> 1 massdataset mutate_ms2() 2024-09-25 20:43:34
#> annotate_metabolites ----------
#> Package Function.used Time
#> 1 metid annotate_metabolites_mass_dataset() 2024-09-25 20:47:07.586516
#> 2 metid annotate_metabolites_mass_dataset() 2024-09-25 20:47:44.92965
#> 3 metid annotate_metabolites_mass_dataset() 2024-09-25 20:49:01.646808
#> merge_mass_dataset ----------
#> Package Function.used Time
#> 1 massdataset merge_mass_dataset 2024-09-25 20:59:42
Trace processing information of object
Then we can use the report_parameters() function to trace processing information of object.
All the analysis results will be placed in a folder named as statistical_analysis.
dir.create(path = "statistical_analysis", showWarnings = FALSE)
report_parameters(object = object, path = "statistical_analysis/")
A html format file named as parameter_report.html will be generated.

Remove redundant metabolites
Remove the redundant annotated metabolites bases on Level and score.
object <-
object %>%
activate_mass_dataset(what = "annotation_table") %>%
group_by(Compound.name) %>%
filter(Level == min(Level)) %>%
filter(SS == max(SS)) %>%
slice_head(n = 1)
object <-
object %>%
activate_mass_dataset(what = "annotation_table") %>%
group_by(variable_id) %>%
filter(Level == min(Level)) %>%
filter(SS == max(SS)) %>%
slice_head(n = 1)
Differential expression metabolites
Calculate the fold changes.
control_sample_id =
object %>%
activate_mass_dataset(what = "sample_info") %>%
filter(group == "Control") %>%
pull(sample_id)
case_sample_id =
object %>%
activate_mass_dataset(what = "sample_info") %>%
filter(group == "Case") %>%
pull(sample_id)
object <-
mutate_fc(object = object,
control_sample_id = control_sample_id,
case_sample_id = case_sample_id,
mean_median = "mean")
#> 110 control samples.
#> 110 case samples.
Calculate p values.
object <-
mutate_p_value(
object = object,
control_sample_id = control_sample_id,
case_sample_id = case_sample_id,
method = "t.test",
p_adjust_methods = "BH"
)
#> 110 control samples.
#> 110 case samples.
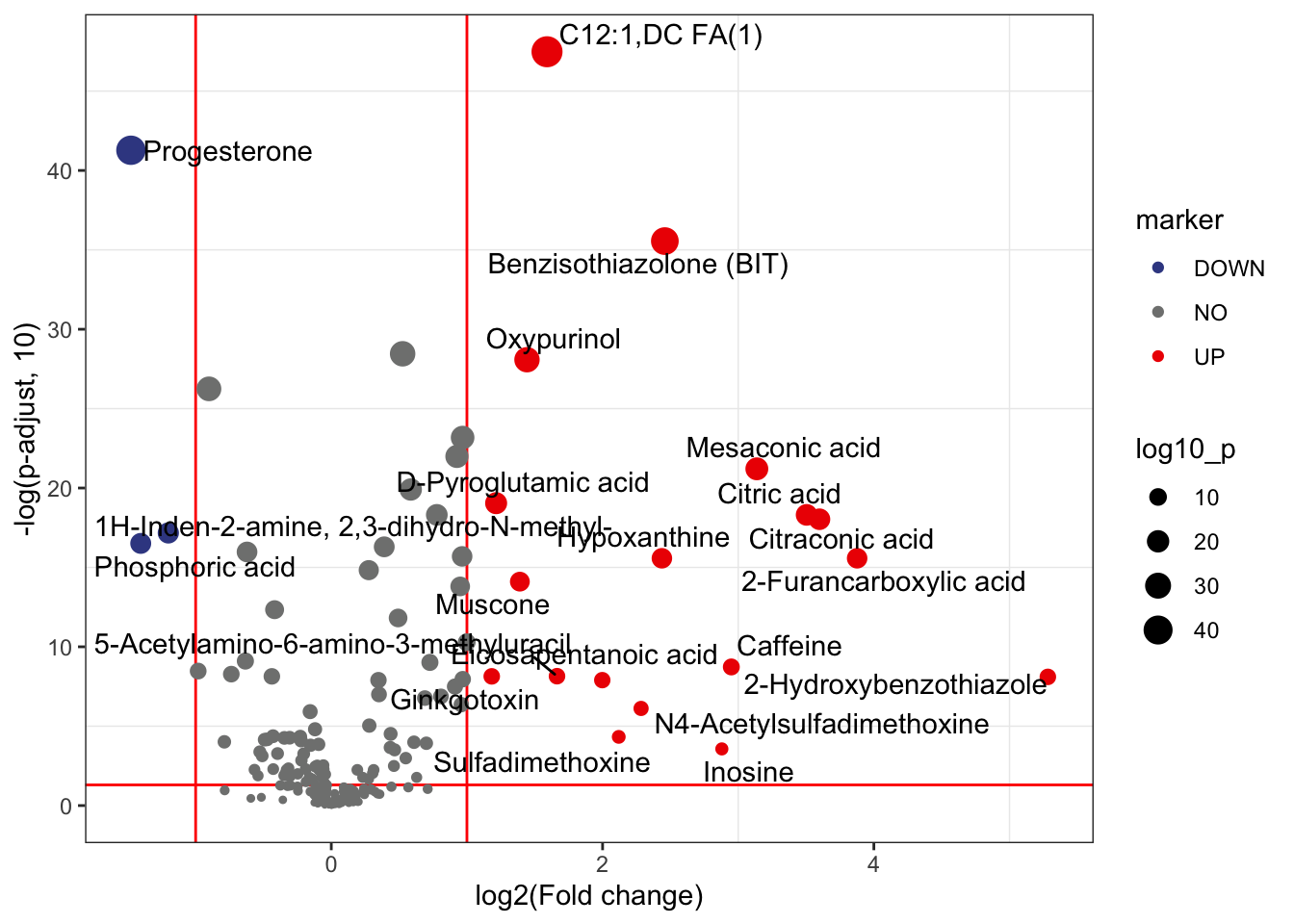
Volcano plot.
volcano_plot(object = object,
add_text = TRUE,
text_from = "Compound.name",
point_size_scale = "p_value") +
scale_size_continuous(range = c(0.5, 5))
Output result
Output the differential expression metabolites.
differential_metabolites <-
extract_variable_info(object = object) %>%
filter(fc > 2 | fc < 0.5) %>%
filter(p_value_adjust < 0.05)
readr::write_csv(differential_metabolites,
file = "statistical_analysis/differential_metabolites.csv")
Save result for subsequent analysis.
object_final <- object
save(object_final, file = "statistical_analysis/object_final")
Session information
sessionInfo()
#> R version 4.4.1 (2024-06-14)
#> Platform: aarch64-apple-darwin20
#> Running under: macOS 15.0
#>
#> Matrix products: default
#> BLAS: /Library/Frameworks/R.framework/Versions/4.4-arm64/Resources/lib/libRblas.0.dylib
#> LAPACK: /Library/Frameworks/R.framework/Versions/4.4-arm64/Resources/lib/libRlapack.dylib; LAPACK version 3.12.0
#>
#> locale:
#> [1] en_US.UTF-8/en_US.UTF-8/en_US.UTF-8/C/en_US.UTF-8/en_US.UTF-8
#>
#> time zone: Asia/Singapore
#> tzcode source: internal
#>
#> attached base packages:
#> [1] grid stats4 stats graphics grDevices utils datasets
#> [8] methods base
#>
#> other attached packages:
#> [1] lubridate_1.9.3 forcats_1.0.0 stringr_1.5.1
#> [4] purrr_1.0.2 readr_2.1.5 tibble_3.2.1
#> [7] tidyverse_2.0.0 metid_1.2.34 metpath_1.0.8
#> [10] ComplexHeatmap_2.20.0 mixOmics_6.28.0 lattice_0.22-6
#> [13] MASS_7.3-61 massstat_1.0.6 tidyr_1.3.1
#> [16] ggfortify_0.4.17 massqc_1.0.7 masscleaner_1.0.12
#> [19] MSnbase_2.30.1 ProtGenerics_1.36.0 S4Vectors_0.42.1
#> [22] Biobase_2.64.0 BiocGenerics_0.50.0 mzR_2.38.0
#> [25] Rcpp_1.0.13 xcms_4.2.3 BiocParallel_1.38.0
#> [28] massprocesser_1.0.10 ggplot2_3.5.1 dplyr_1.1.4
#> [31] magrittr_2.0.3 masstools_1.0.13 massdataset_1.0.34
#> [34] tidymass_1.0.9
#>
#> loaded via a namespace (and not attached):
#> [1] fs_1.6.4 matrixStats_1.3.0
#> [3] bitops_1.0-8 fit.models_0.64
#> [5] httr_1.4.7 RColorBrewer_1.1-3
#> [7] doParallel_1.0.17 tools_4.4.1
#> [9] doRNG_1.8.6 backports_1.5.0
#> [11] utf8_1.2.4 R6_2.5.1
#> [13] lazyeval_0.2.2 GetoptLong_1.0.5
#> [15] withr_3.0.1 prettyunits_1.2.0
#> [17] gridExtra_2.3 preprocessCore_1.66.0
#> [19] cli_3.6.3 fastDummies_1.7.4
#> [21] labeling_0.4.3 sass_0.4.9
#> [23] mvtnorm_1.3-1 robustbase_0.99-4
#> [25] randomForest_4.7-1.1 proxy_0.4-27
#> [27] pbapply_1.7-2 foreign_0.8-87
#> [29] rrcov_1.7-6 MetaboCoreUtils_1.12.0
#> [31] parallelly_1.38.0 itertools_0.1-3
#> [33] limma_3.60.4 readxl_1.4.3
#> [35] rstudioapi_0.16.0 impute_1.78.0
#> [37] generics_0.1.3 shape_1.4.6.1
#> [39] vroom_1.6.5 zip_2.3.1
#> [41] Matrix_1.7-0 MALDIquant_1.22.3
#> [43] fansi_1.0.6 abind_1.4-5
#> [45] lifecycle_1.0.4 yaml_2.3.10
#> [47] SummarizedExperiment_1.34.0 SparseArray_1.4.8
#> [49] crayon_1.5.3 PSMatch_1.8.0
#> [51] KEGGREST_1.44.1 pillar_1.9.0
#> [53] knitr_1.48 GenomicRanges_1.56.1
#> [55] rjson_0.2.22 corpcor_1.6.10
#> [57] codetools_0.2-20 glue_1.7.0
#> [59] pcaMethods_1.96.0 data.table_1.16.0
#> [61] remotes_2.5.0 MultiAssayExperiment_1.30.3
#> [63] vctrs_0.6.5 png_0.1-8
#> [65] cellranger_1.1.0 gtable_0.3.5
#> [67] cachem_1.1.0 xfun_0.47
#> [69] openxlsx_4.2.7 S4Arrays_1.4.1
#> [71] tidygraph_1.3.1 pcaPP_2.0-5
#> [73] ncdf4_1.23 iterators_1.0.14
#> [75] statmod_1.5.0 bit64_4.0.5
#> [77] robust_0.7-5 progress_1.2.3
#> [79] GenomeInfoDb_1.40.1 rprojroot_2.0.4
#> [81] bslib_0.8.0 affyio_1.74.0
#> [83] rpart_4.1.23 colorspace_2.1-1
#> [85] DBI_1.2.3 Hmisc_5.1-3
#> [87] nnet_7.3-19 tidyselect_1.2.1
#> [89] bit_4.0.5 compiler_4.4.1
#> [91] MassSpecWavelet_1.70.0 htmlTable_2.4.3
#> [93] DelayedArray_0.30.1 plotly_4.10.4
#> [95] bookdown_0.40 checkmate_2.3.2
#> [97] scales_1.3.0 DEoptimR_1.1-3
#> [99] affy_1.82.0 digest_0.6.37
#> [101] rmarkdown_2.28 XVector_0.44.0
#> [103] htmltools_0.5.8.1 pkgconfig_2.0.3
#> [105] base64enc_0.1-3 MatrixGenerics_1.16.0
#> [107] highr_0.11 fastmap_1.2.0
#> [109] rlang_1.1.4 GlobalOptions_0.1.2
#> [111] htmlwidgets_1.6.4 UCSC.utils_1.0.0
#> [113] farver_2.1.2 jquerylib_0.1.4
#> [115] jsonlite_1.8.8 MsExperiment_1.6.0
#> [117] mzID_1.42.0 RCurl_1.98-1.16
#> [119] Formula_1.2-5 GenomeInfoDbData_1.2.12
#> [121] patchwork_1.2.0 munsell_0.5.1
#> [123] viridis_0.6.5 MsCoreUtils_1.16.1
#> [125] vsn_3.72.0 furrr_0.3.1
#> [127] stringi_1.8.4 ggraph_2.2.1
#> [129] zlibbioc_1.50.0 plyr_1.8.9
#> [131] parallel_4.4.1 listenv_0.9.1
#> [133] ggrepel_0.9.5 Biostrings_2.72.1
#> [135] MsFeatures_1.12.0 graphlayouts_1.1.1
#> [137] hms_1.1.3 Spectra_1.14.1
#> [139] circlize_0.4.16 igraph_2.0.3
#> [141] QFeatures_1.14.2 rngtools_1.5.2
#> [143] reshape2_1.4.4 XML_3.99-0.17
#> [145] evaluate_0.24.0 blogdown_1.19
#> [147] BiocManager_1.30.25 tzdb_0.4.0
#> [149] foreach_1.5.2 missForest_1.5
#> [151] tweenr_2.0.3 polyclip_1.10-7
#> [153] future_1.34.0 clue_0.3-65
#> [155] ggforce_0.4.2 AnnotationFilter_1.28.0
#> [157] e1071_1.7-14 RSpectra_0.16-2
#> [159] ggcorrplot_0.1.4.1 viridisLite_0.4.2
#> [161] class_7.3-22 rARPACK_0.11-0
#> [163] memoise_2.0.1 ellipse_0.5.0
#> [165] IRanges_2.38.1 cluster_2.1.6
#> [167] timechange_0.3.0 globals_0.16.3
#> [169] here_1.0.1