Data preparation
In mass_dataset class object, it contains all the processing information in it. We can trace the analysis and parameters so we can do the reproducible analysis.
library(massdataset)
data("expression_data")
data("sample_info")
data("variable_info")
object =
create_mass_dataset(
expression_data = expression_data,
sample_info = sample_info,
variable_info = variable_info
)
library(tidyverse)
object =
object %>%
activate_mass_dataset(what = "expression_data") %>%
filter(!is.na(QC_1))
object =
object %>%
activate_mass_dataset(what = "expression_data") %>%
filter(!is.na(QC_2))
object =
object %>%
mutate_mean_intensity()
object =
object %>%
mutate_median_intensity() %>%
mutate_rsd()
process_info
process_info = extract_process_info(object)
process_info
#> $create_mass_dataset
#> --------------------
#> pacakge_name: massdataset
#> function_name: create_mass_dataset()
#> time: 2024-09-25 19:11:39.449469
#> parameters:
#> no : no
#>
#> $filter
#> $filter[[1]]
#> --------------------
#> pacakge_name: massdataset
#> function_name: filter()
#> time: 2024-09-25 19:11:39.843294
#> parameters:
#> parameter : `~!is.na(QC_1)`
#>
#> $filter[[2]]
#> --------------------
#> pacakge_name: massdataset
#> function_name: filter()
#> time: 2024-09-25 19:11:39.844175
#> parameters:
#> parameter : `~!is.na(QC_2)`
#>
#>
#> $mutate_mean_intensity
#> --------------------
#> pacakge_name: massdataset
#> function_name: mutate_mean_intensity()
#> time: 2024-09-25 19:11:39.847539
#> parameters:
#> according_to_samples : c("Blank_3", "Blank_4", "QC_1", "QC_2", "PS4P1", "PS4P2", "PS4P3", "PS4P4")
#>
#> $mutate_median_intensity
#> --------------------
#> pacakge_name: massdataset
#> function_name: mutate_median_intensity()
#> time: 2024-09-25 19:11:39.854193
#> parameters:
#> according_to_samples : c("Blank_3", "Blank_4", "QC_1", "QC_2", "PS4P1", "PS4P2", "PS4P3", "PS4P4")
#>
#> $mutate_rsd
#> --------------------
#> pacakge_name: massdataset
#> function_name: mutate_rsd()
#> time: 2024-09-25 19:11:39.85757
#> parameters:
#> according_to_samples : c("Blank_3", "Blank_4", "QC_1", "QC_2", "PS4P1", "PS4P2", "PS4P3", "PS4P4")
The process_info contains all the steps which are ordered by time.
process_info$mutate_median_intensity
#> --------------------
#> pacakge_name: massdataset
#> function_name: mutate_median_intensity()
#> time: 2024-09-25 19:11:39.854193
#> parameters:
#> according_to_samples : c("Blank_3", "Blank_4", "QC_1", "QC_2", "PS4P1", "PS4P2", "PS4P3", "PS4P4")
process_info$mutate_median_intensity@parameter
#> $according_to_samples
#> [1] "Blank_3" "Blank_4" "QC_1" "QC_2" "PS4P1" "PS4P2" "PS4P3"
#> [8] "PS4P4"
Output html processing information
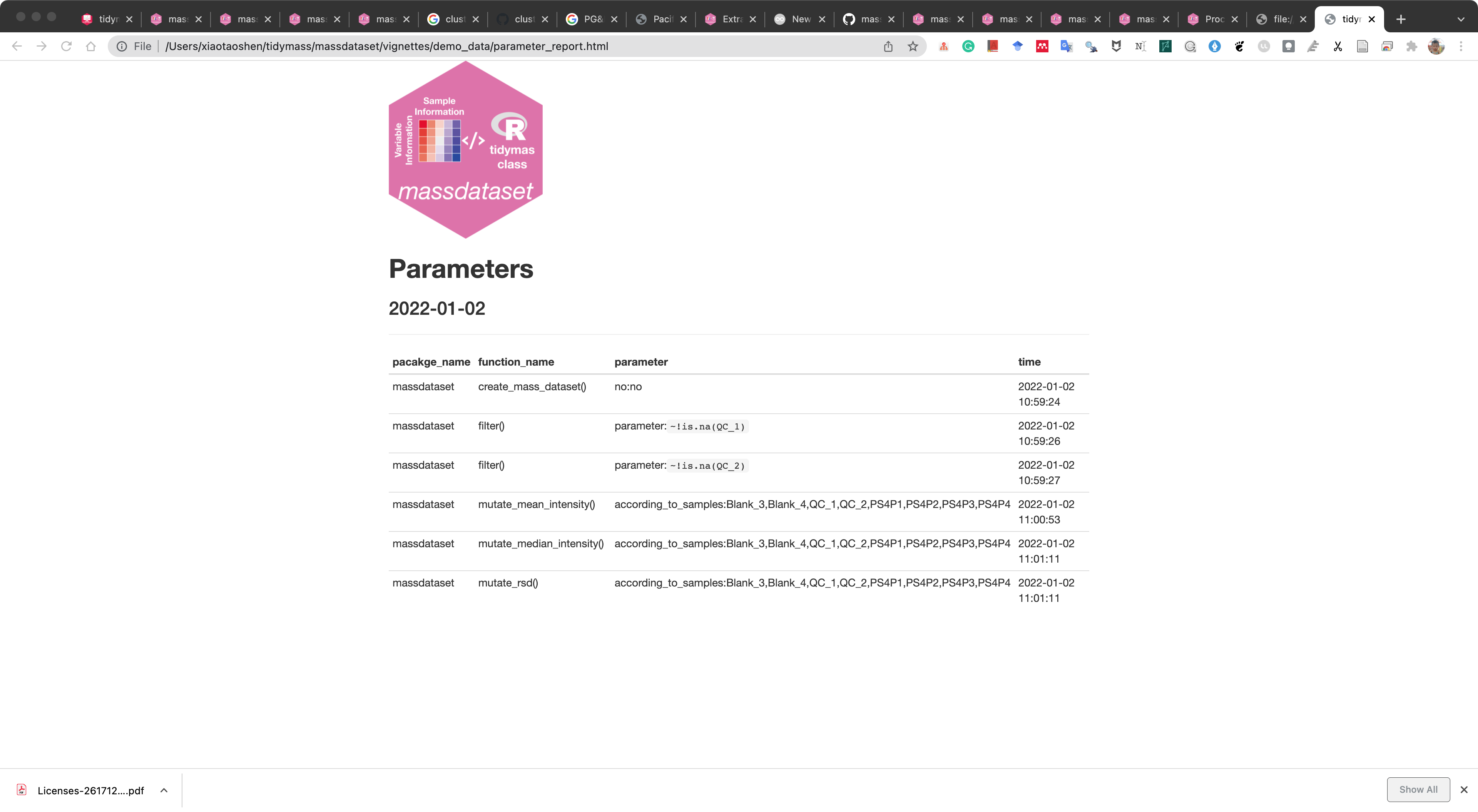
We can output the process_info into a html format file, so we can know what processing steps have been made to this object and the accurate parameters.
Then we can use report_parameters() to output this into a html file.
report_parameters(object = object,
path = "data_cleaning")
A html file named as parameter_report.html will be generated and saved in data_cleaning folder.
Session information
sessionInfo()
#> R version 4.4.1 (2024-06-14)
#> Platform: aarch64-apple-darwin20
#> Running under: macOS 15.0
#>
#> Matrix products: default
#> BLAS: /Library/Frameworks/R.framework/Versions/4.4-arm64/Resources/lib/libRblas.0.dylib
#> LAPACK: /Library/Frameworks/R.framework/Versions/4.4-arm64/Resources/lib/libRlapack.dylib; LAPACK version 3.12.0
#>
#> locale:
#> [1] en_US.UTF-8/en_US.UTF-8/en_US.UTF-8/C/en_US.UTF-8/en_US.UTF-8
#>
#> time zone: Asia/Singapore
#> tzcode source: internal
#>
#> attached base packages:
#> [1] stats graphics grDevices utils datasets methods base
#>
#> other attached packages:
#> [1] lubridate_1.9.3 forcats_1.0.0 stringr_1.5.1 purrr_1.0.2
#> [5] readr_2.1.5 tidyr_1.3.1 tibble_3.2.1 tidyverse_2.0.0
#> [9] ggplot2_3.5.1 dplyr_1.1.4 magrittr_2.0.3 masstools_1.0.13
#> [13] massdataset_1.0.34
#>
#> loaded via a namespace (and not attached):
#> [1] bitops_1.0-8 pbapply_1.7-2
#> [3] remotes_2.5.0 rlang_1.1.4
#> [5] clue_0.3-65 GetoptLong_1.0.5
#> [7] matrixStats_1.3.0 compiler_4.4.1
#> [9] png_0.1-8 vctrs_0.6.5
#> [11] reshape2_1.4.4 ProtGenerics_1.36.0
#> [13] pkgconfig_2.0.3 shape_1.4.6.1
#> [15] crayon_1.5.3 fastmap_1.2.0
#> [17] XVector_0.44.0 utf8_1.2.4
#> [19] rmarkdown_2.28 tzdb_0.4.0
#> [21] preprocessCore_1.66.0 UCSC.utils_1.0.0
#> [23] xfun_0.47 MultiAssayExperiment_1.30.3
#> [25] zlibbioc_1.50.0 cachem_1.1.0
#> [27] GenomeInfoDb_1.40.1 jsonlite_1.8.8
#> [29] DelayedArray_0.30.1 BiocParallel_1.38.0
#> [31] parallel_4.4.1 cluster_2.1.6
#> [33] R6_2.5.1 bslib_0.8.0
#> [35] stringi_1.8.4 RColorBrewer_1.1-3
#> [37] limma_3.60.4 GenomicRanges_1.56.1
#> [39] jquerylib_0.1.4 Rcpp_1.0.13
#> [41] bookdown_0.40 SummarizedExperiment_1.34.0
#> [43] iterators_1.0.14 knitr_1.48
#> [45] IRanges_2.38.1 timechange_0.3.0
#> [47] Matrix_1.7-0 igraph_2.0.3
#> [49] tidyselect_1.2.1 rstudioapi_0.16.0
#> [51] abind_1.4-5 yaml_2.3.10
#> [53] affy_1.82.0 doParallel_1.0.17
#> [55] codetools_0.2-20 blogdown_1.19
#> [57] lattice_0.22-6 plyr_1.8.9
#> [59] withr_3.0.1 Biobase_2.64.0
#> [61] evaluate_0.24.0 zip_2.3.1
#> [63] circlize_0.4.16 BiocManager_1.30.25
#> [65] affyio_1.74.0 pillar_1.9.0
#> [67] MatrixGenerics_1.16.0 foreach_1.5.2
#> [69] stats4_4.4.1 MSnbase_2.30.1
#> [71] MALDIquant_1.22.3 ncdf4_1.23
#> [73] generics_0.1.3 RCurl_1.98-1.16
#> [75] rprojroot_2.0.4 hms_1.1.3
#> [77] S4Vectors_0.42.1 munsell_0.5.1
#> [79] scales_1.3.0 glue_1.7.0
#> [81] lazyeval_0.2.2 tools_4.4.1
#> [83] mzID_1.42.0 QFeatures_1.14.2
#> [85] vsn_3.72.0 mzR_2.38.0
#> [87] openxlsx_4.2.7 XML_3.99-0.17
#> [89] grid_4.4.1 impute_1.78.0
#> [91] MsCoreUtils_1.16.1 colorspace_2.1-1
#> [93] GenomeInfoDbData_1.2.12 PSMatch_1.8.0
#> [95] cli_3.6.3 fansi_1.0.6
#> [97] S4Arrays_1.4.1 ComplexHeatmap_2.20.0
#> [99] AnnotationFilter_1.28.0 pcaMethods_1.96.0
#> [101] gtable_0.3.5 sass_0.4.9
#> [103] digest_0.6.37 BiocGenerics_0.50.0
#> [105] SparseArray_1.4.8 rjson_0.2.22
#> [107] htmltools_0.5.8.1 lifecycle_1.0.4
#> [109] httr_1.4.7 here_1.0.1
#> [111] statmod_1.5.0 GlobalOptions_0.1.2
#> [113] MASS_7.3-61