
Merge two `mass_dataset`
Xiaotao Shen
Created on 2021-12-04 and updated on 2026-03-04
Source:vignettes/merge_two_mass_dataset.Rmd
merge_two_mass_dataset.RmdIn massdataset package, the
merge_mass_dataset is more powerful to merge tow
mass_dataset class objects.
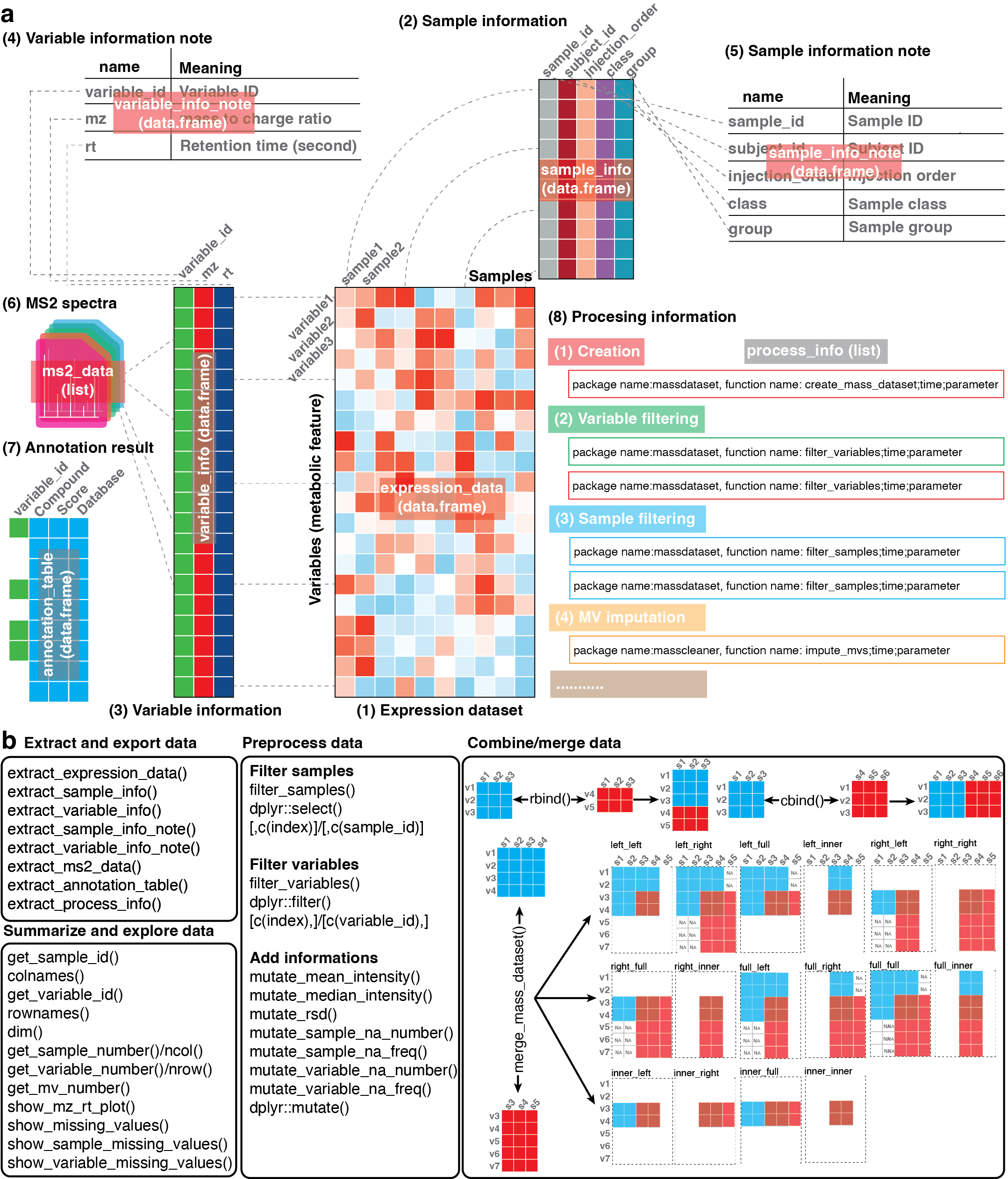
Diagram showing overlap between two mass_dataset
objects before merging.
Data preparation
library(massdataset)
library(tidyverse)
data("expression_data")
data("sample_info")
data("variable_info")
object =
create_mass_dataset(
expression_data = expression_data,
sample_info = sample_info,
variable_info = variable_info
)
object1 = object[1:10, 1:5]
object2 = object[5:15, 3:7]
colnames(object1)
#> [1] "Blank_3" "Blank_4" "QC_1" "QC_2" "PS4P1"
colnames(object2)
#> [1] "QC_1" "QC_2" "PS4P1" "PS4P2" "PS4P3"
rownames(object1)
#> [1] "M136T55_2_POS" "M79T35_POS" "M307T548_POS" "M183T224_POS"
#> [5] "M349T47_POS" "M182T828_POS" "M299T359_POS" "M348T844_POS"
#> [9] "M344T471_POS" "M181T436_POS"
rownames(object2)
#> [1] "M349T47_POS" "M182T828_POS" "M299T359_POS" "M348T844_POS" "M344T471_POS"
#> [6] "M181T436_POS" "M345T195_POS" "M304T825_POS" "M137T196_POS" "M359T638_POS"
#> [11] "M270T507_POS"
Merge two mass_dataset
object =
merge_mass_dataset(x = object1, y = object2,
sample_direction = "left",
variable_direction = "left",
sample_by = "sample_id",
variable_by = "variable_id")
extract_expression_data(object1)
#> Blank_3 Blank_4 QC_1 QC_2 PS4P1
#> M136T55_2_POS NA NA 1857924.8 1037763.8 1494436.1
#> M79T35_POS NA NA 2821550.2 1304875.3 2471336.1
#> M307T548_POS NA NA 410387.6 273687.8 288590.2
#> M183T224_POS NA NA NA NA NA
#> M349T47_POS NA NA 8730104.8 4105598.5 5141073.2
#> M182T828_POS 3761892.6 2572593.4 NA 3662819.1 5700534.8
#> M299T359_POS NA NA 3688690.6 2892719.6 1401632.7
#> M348T844_POS NA NA NA 3131157.7 NA
#> M344T471_POS NA NA 589957.0 408610.8 NA
#> M181T436_POS 249352.6 131374.5 248764.1 208789.4 423991.8
extract_expression_data(object2)
#> QC_1 QC_2 PS4P1 PS4P2 PS4P3
#> M349T47_POS 8730104.8 4105598.5 5141073.2 8424315.6 7896633.3
#> M182T828_POS NA 3662819.1 5700534.8 4600172.4 5557014.6
#> M299T359_POS 3688690.6 2892719.6 1401632.7 4055989.5 1577496.3
#> M348T844_POS NA 3131157.7 NA NA 3643606.4
#> M344T471_POS 589957.0 408610.8 NA 276913.4 304611.5
#> M181T436_POS 248764.1 208789.4 423991.8 449021.6 357037.7
#> M345T195_POS NA 5776921.1 NA NA NA
#> M304T825_POS 2816826.6 237776.3 439981.6 510661.8 415109.5
#> M137T196_POS NA 11028014.0 NA NA NA
#> M359T638_POS 1367524.9 1044288.4 1786016.1 1878777.4 1025039.9
#> M270T507_POS 107442.7 NA NA NA 60286.3
extract_expression_data(object)
#> Blank_3 Blank_4 QC_1 QC_2 PS4P1
#> M136T55_2_POS NA NA 1857924.8 1037763.8 1494436.1
#> M79T35_POS NA NA 2821550.2 1304875.3 2471336.1
#> M307T548_POS NA NA 410387.6 273687.8 288590.2
#> M183T224_POS NA NA NA NA NA
#> M349T47_POS NA NA 8730104.8 4105598.5 5141073.2
#> M182T828_POS 3761892.6 2572593.4 NA 3662819.1 5700534.8
#> M299T359_POS NA NA 3688690.6 2892719.6 1401632.7
#> M348T844_POS NA NA NA 3131157.7 NA
#> M344T471_POS NA NA 589957.0 408610.8 NA
#> M181T436_POS 249352.6 131374.5 248764.1 208789.4 423991.8Session information
sessionInfo()
#> R version 4.5.2 (2025-10-31)
#> Platform: aarch64-apple-darwin20
#> Running under: macOS Tahoe 26.3
#>
#> Matrix products: default
#> BLAS: /System/Library/Frameworks/Accelerate.framework/Versions/A/Frameworks/vecLib.framework/Versions/A/libBLAS.dylib
#> LAPACK: /Library/Frameworks/R.framework/Versions/4.5-arm64/Resources/lib/libRlapack.dylib; LAPACK version 3.12.1
#>
#> locale:
#> [1] en_US.UTF-8/en_US.UTF-8/en_US.UTF-8/C/en_US.UTF-8/en_US.UTF-8
#>
#> time zone: Asia/Singapore
#> tzcode source: internal
#>
#> attached base packages:
#> [1] stats graphics grDevices utils datasets methods base
#>
#> other attached packages:
#> [1] lubridate_1.9.4 forcats_1.0.0 stringr_1.5.1 purrr_1.1.0
#> [5] readr_2.1.5 tidyr_1.3.1 tibble_3.3.0 tidyverse_2.0.0
#> [9] magrittr_2.0.3 dplyr_1.1.4 ggplot2_4.0.2 massdataset_0.99.3
#>
#> loaded via a namespace (and not attached):
#> [1] tidyselect_1.2.1 farver_2.1.2
#> [3] S7_0.2.0 fastmap_1.2.0
#> [5] digest_0.6.37 timechange_0.3.0
#> [7] lifecycle_1.0.4 cluster_2.1.8.1
#> [9] compiler_4.5.2 rlang_1.1.6
#> [11] sass_0.4.10 tools_4.5.2
#> [13] yaml_2.3.10 knitr_1.50
#> [15] S4Arrays_1.8.1 htmlwidgets_1.6.4
#> [17] DelayedArray_0.34.1 RColorBrewer_1.1-3
#> [19] abind_1.4-8 withr_3.0.2
#> [21] BiocGenerics_0.54.0 desc_1.4.3
#> [23] grid_4.5.2 stats4_4.5.2
#> [25] colorspace_2.1-1 scales_1.4.0
#> [27] iterators_1.0.14 dichromat_2.0-0.1
#> [29] SummarizedExperiment_1.38.1 cli_3.6.5
#> [31] rmarkdown_2.29 crayon_1.5.3
#> [33] ragg_1.4.0 generics_0.1.4
#> [35] rstudioapi_0.17.1 httr_1.4.7
#> [37] tzdb_0.5.0 rjson_0.2.23
#> [39] cachem_1.1.0 parallel_4.5.2
#> [41] XVector_0.48.0 matrixStats_1.5.0
#> [43] vctrs_0.6.5 Matrix_1.7-4
#> [45] jsonlite_2.0.0 IRanges_2.42.0
#> [47] hms_1.1.3 GetoptLong_1.0.5
#> [49] S4Vectors_0.48.0 clue_0.3-66
#> [51] systemfonts_1.2.3 foreach_1.5.2
#> [53] jquerylib_0.1.4 glue_1.8.0
#> [55] pkgdown_2.1.3 codetools_0.2-20
#> [57] stringi_1.8.7 shape_1.4.6.1
#> [59] gtable_0.3.6 GenomeInfoDb_1.44.2
#> [61] GenomicRanges_1.60.0 UCSC.utils_1.4.0
#> [63] ComplexHeatmap_2.24.1 pillar_1.11.0
#> [65] htmltools_0.5.8.1 GenomeInfoDbData_1.2.14
#> [67] circlize_0.4.16 R6_2.6.1
#> [69] textshaping_1.0.1 doParallel_1.0.17
#> [71] evaluate_1.0.4 Biobase_2.68.0
#> [73] lattice_0.22-7 png_0.1-8
#> [75] openxlsx_4.2.8 bslib_0.9.0
#> [77] Rcpp_1.1.0 zip_2.3.3
#> [79] SparseArray_1.8.1 xfun_0.53
#> [81] fs_1.6.6 MatrixGenerics_1.20.0
#> [83] pkgconfig_2.0.3 GlobalOptions_0.1.2